Preview
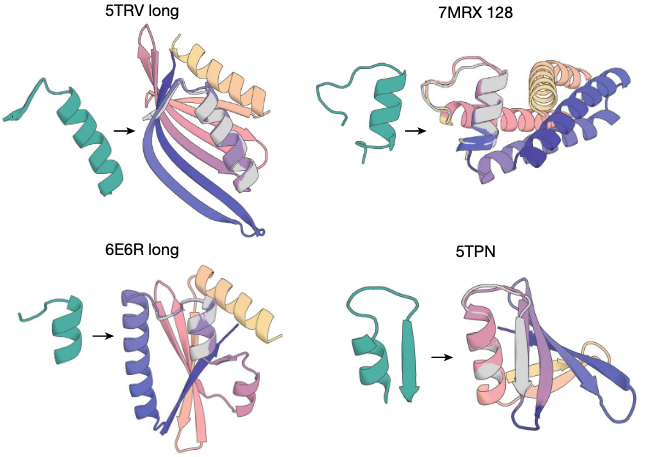
Generating (designing) protein binder. Various applications of RFdiffusion
How can we learn on protein structures
Challenges of proteins vs Images
- Strong geometric constraints (4 backbone heavy atoms, 3 covalent bonds, continuous chain)
- Must also have a sequence that can encode it
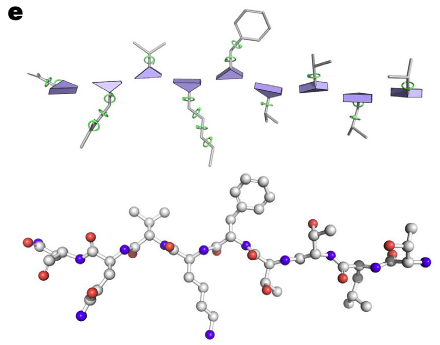 (Jumper et al., 2020;AF2)
(Jumper et al., 2020;AF2) 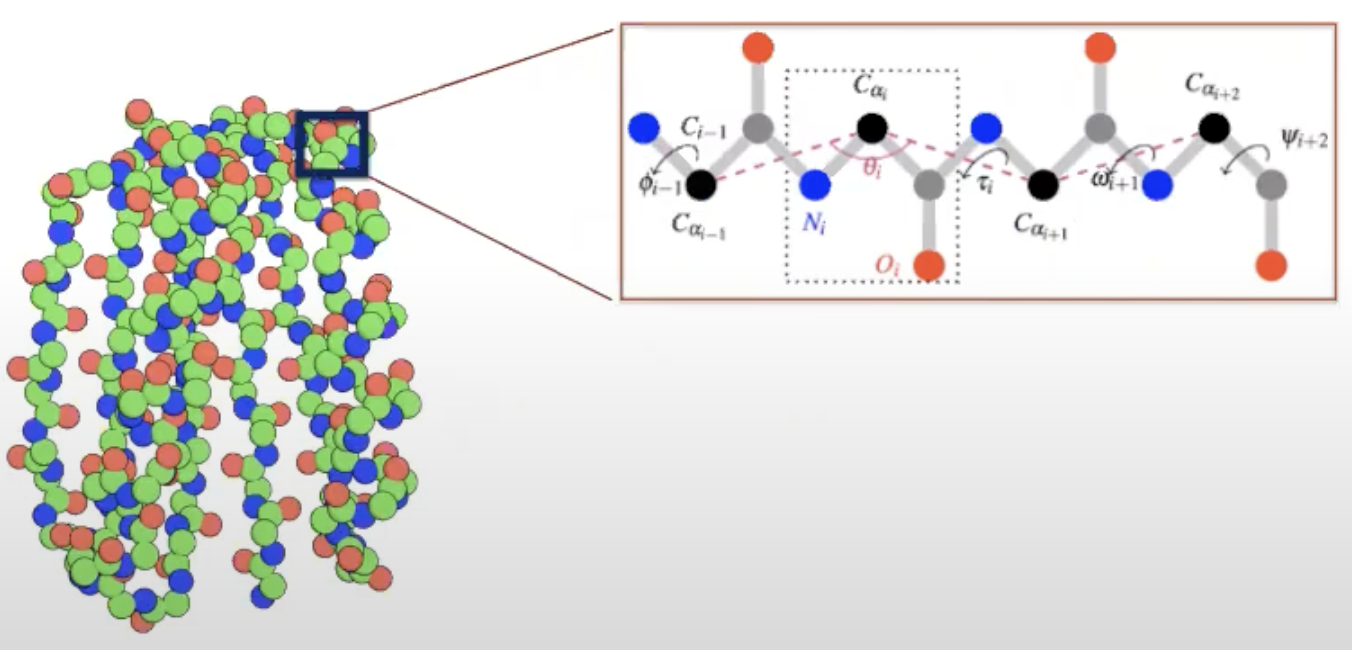
Challenges of proteins vs Images
How to represent a protein backbone
- atom level : too complicated
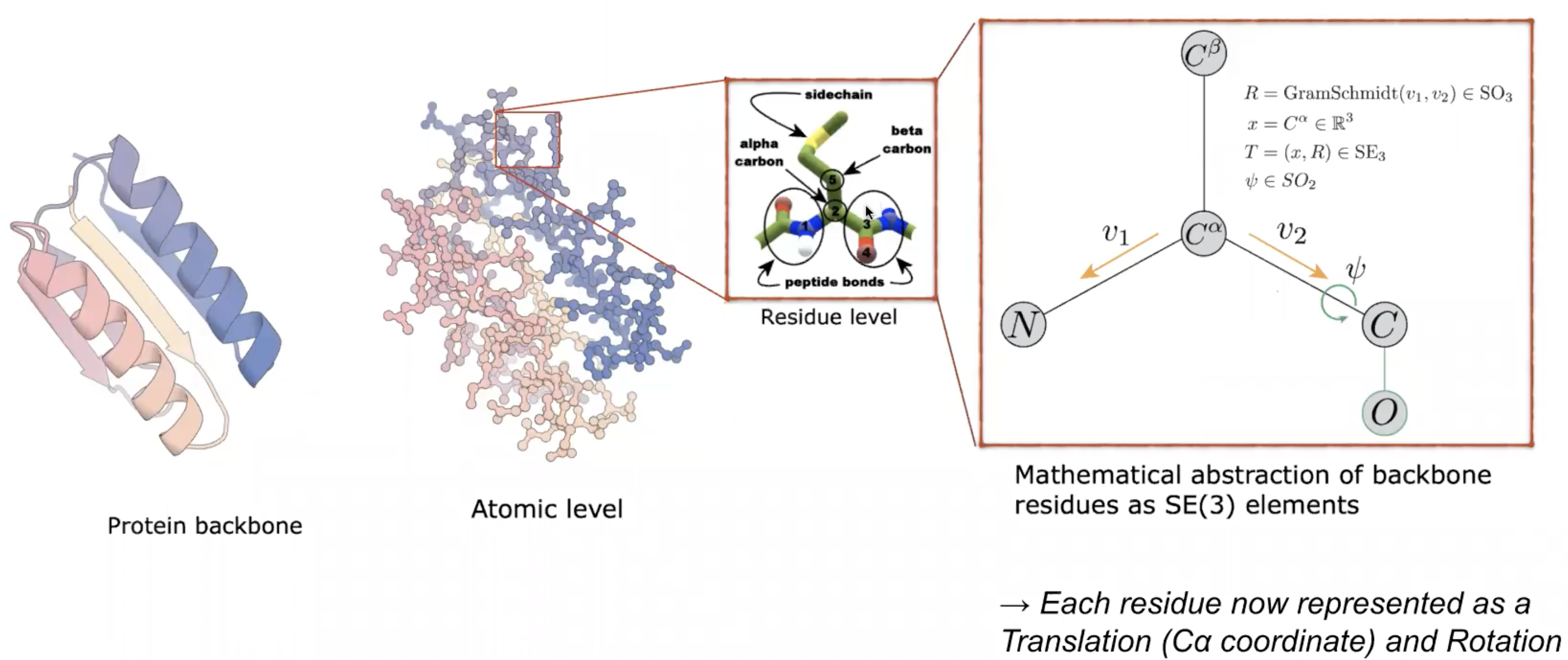
Challenges of proteins vs Images
Diffusion models as an attrative framework for protein design
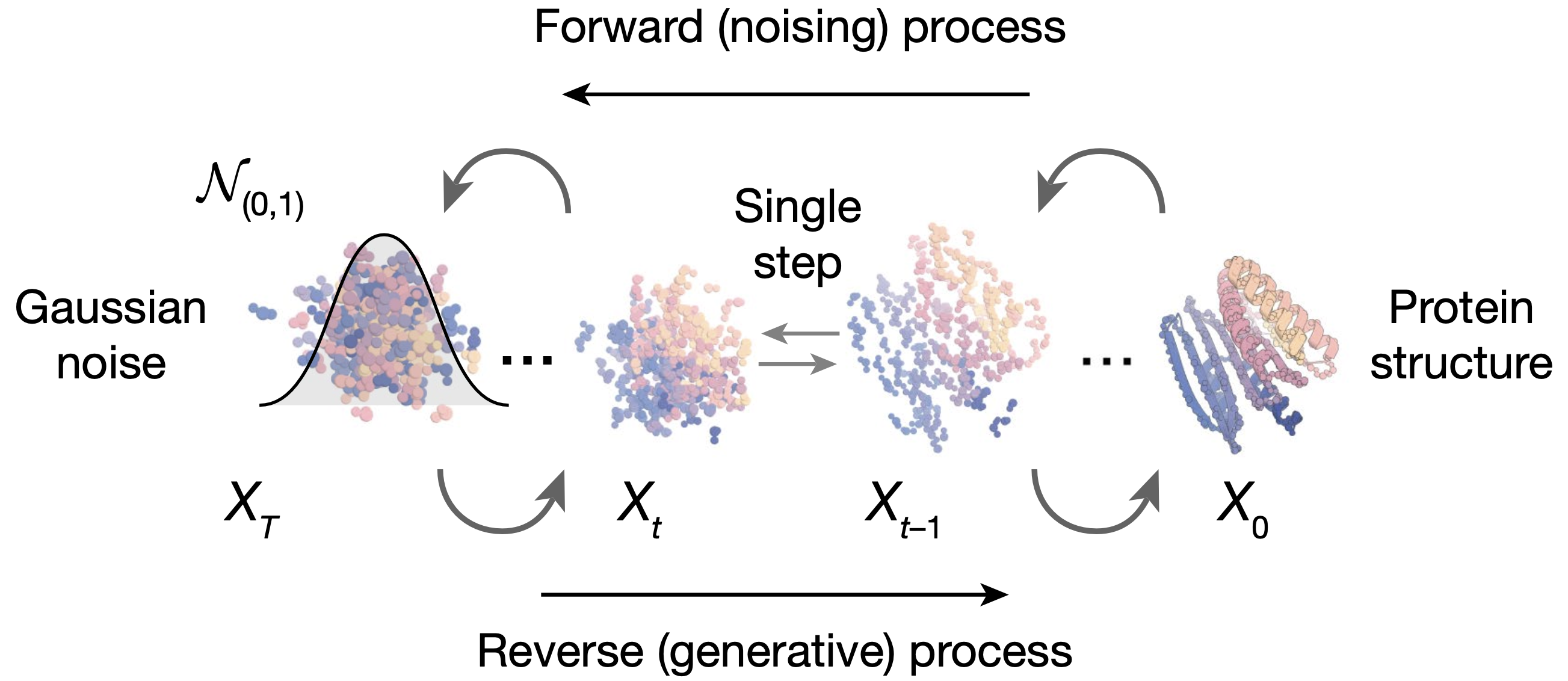
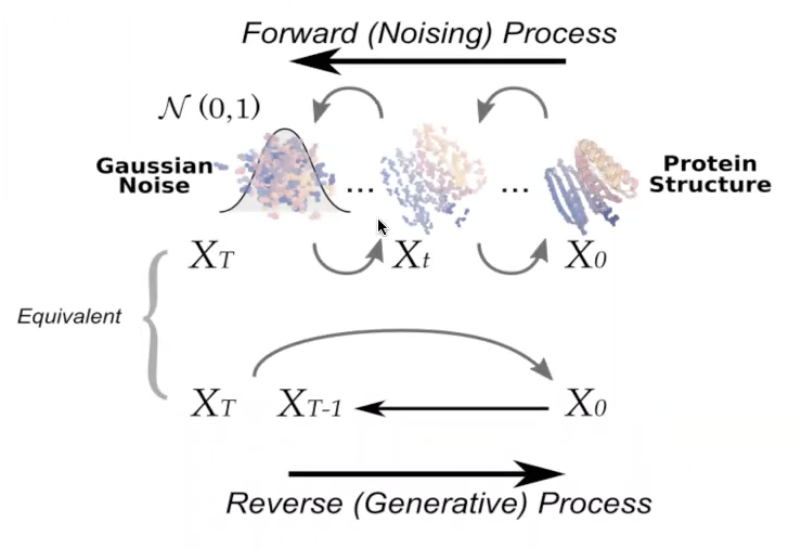
RoseTTA Fold (author's previous work) can be easily used without much changes.
Training summary
- Dataset: PDB (384 amino acids; i.e. no cropping), clustered by sequence similairty
- 200 time steps (t=0: true structures, t=200:
3D Gaussianuniformframe rotations) - Self-conditioning of predicted
- Model was initialised from RoseTTAFold structure prediction weights
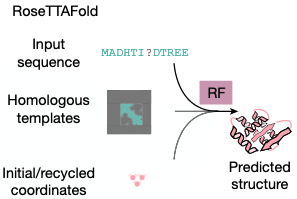
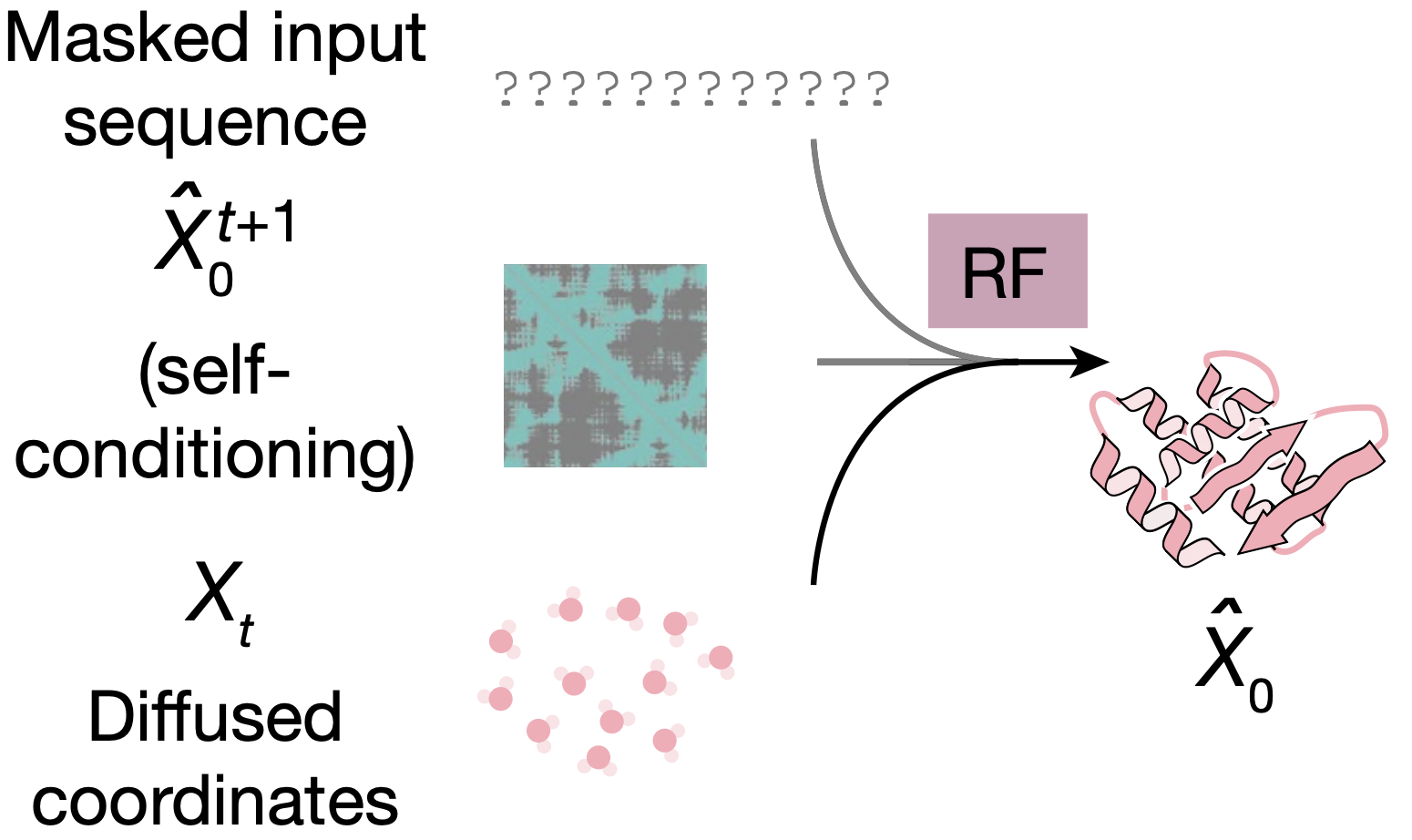
Main inputs: Input sequence vs. Diffused coordinates
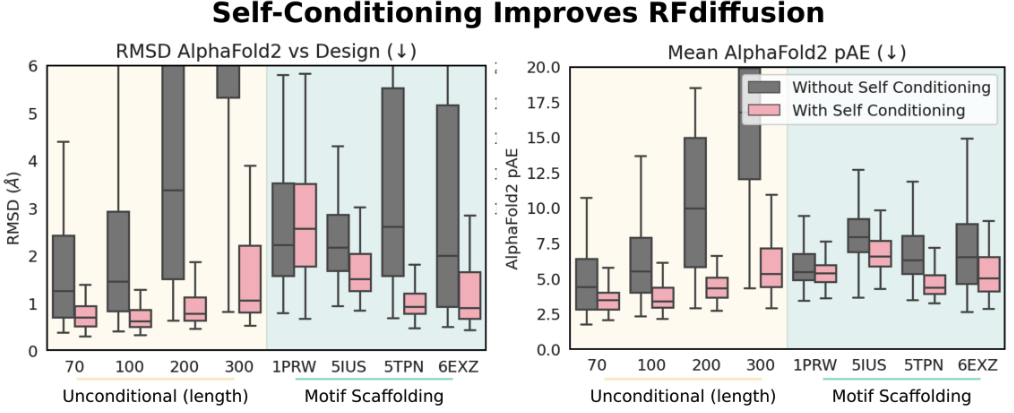
Allowing the model to 'self-condition' improves RFdiffusion
-
In RFdiffusion, the model receives its previous prediction as a template input (‘self-conditioning’).
-
At each timestep
-
The next coordinate input to the model (

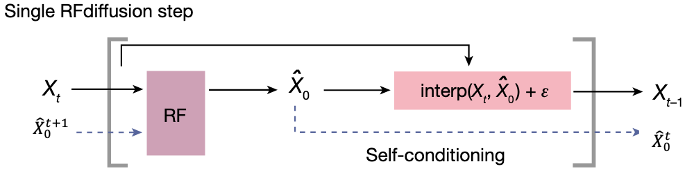
Allowing the model to 'self-condition' improves RFdiffusion
Generate B.B.
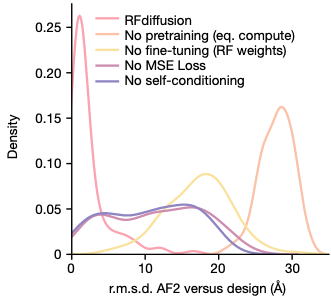
Training from pre-trained RoseTTAFold weights makes training computationally tractable
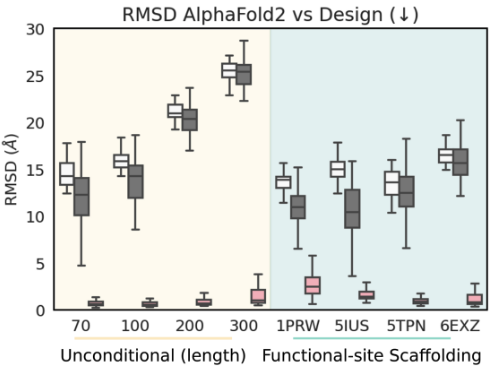
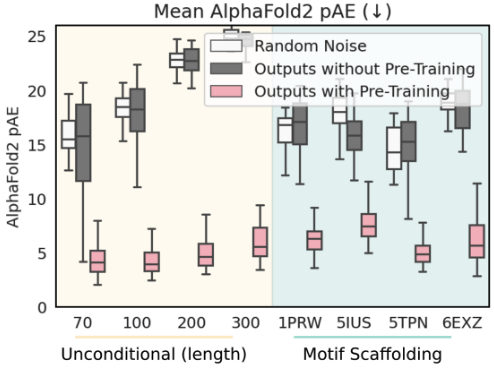
Ablation study
performance: RF diffusion >> MSE Loss
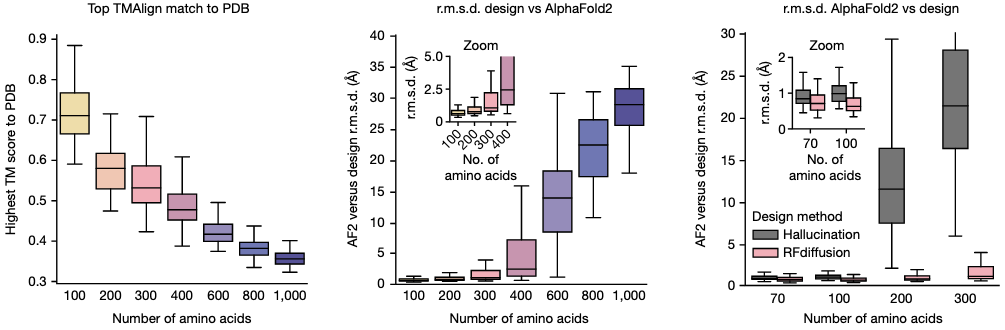
Strengths
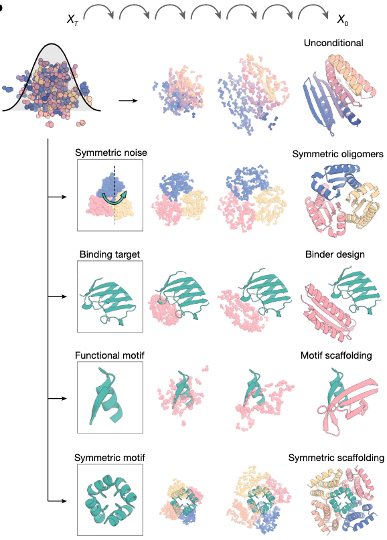
Unconditional generation
Discussion
- How to evaluate the designed proteins
- Directly evaluated the designed proteins with experiments
- Is the RFdiffusion novel method?
- my thought: More impact on the evaluatio with experiments
- What are the other factors affect to designing performance
Conclusion
- There are various use cases of protein design model
- Finetunning generative model with pre-trained structual prediction models (or simulations)
TIL
- Protein can be represented by
Recommendations
Presentation of main authors
Q&A
References
- Watson, Joseph L., David Juergens, Nathaniel R. Bennett, Brian L. Trippe, Jason Yim, Helen E. Eisenach, Woody Ahern, et al. 2023. “De Novo Design of Protein Structure and Function with RFdiffusion.” Nature 620 (7976): 1089–1100. https://doi.org/10.1038/s41586-023-06415-8.
- Joe Watson & David Juergens, presented at Tuesday February 14th, 4-5 pm EST, RFDiffusion: Accurate protein design using structure prediction and diffusion generative models, https://www.youtube.com/watch?v=wIHwHDt2NoI, accessed at 2025.03.17
- Baek, Minkyung, Frank DiMaio, Ivan Anishchenko, Justas Dauparas, Sergey Ovchinnikov, Gyu Rie Lee, Jue Wang, et al. 2021. “Accurate Prediction of Protein Structures and Interactions Using a Three-Track Neural Network.” Science 373 (6557): 871–76. https://doi.org/10.1126/science.abj8754.
- Dauparas, J., I. Anishchenko, N. Bennett, H. Bai, R. J. Ragotte, L. F. Milles, B. I. M. Wicky, et al. 2022. “Robust Deep Learning–Based Protein Sequence Design Using ProteinMPNN.” Science 378 (6615): 49–56. https://doi.org/10.1126/science.add2187.
- Jumper, John, Richard Evans, Alexander Pritzel, Tim Green, Michael Figurnov, Olaf Ronneberger, Kathryn Tunyasuvunakool, et al. 2021. “Highly Accurate Protein Structure Prediction with AlphaFold.” Nature 596 (7873): 583–89. https://doi.org/10.1038/s41586-021-03819-2.
Supplementary
motif scaffolding